Cross linking
Content
This guide is based on and cites the preprint A Research Software Engineering Workflow for Computational Science and Engineering (Maric, Gläser, Lehr et al.; see the literature).
Goal
When you publish, you should ensure that all the relevant data (including the code) can be found. To do so, connect your published evaluation data, code and papers with PIDs and git tags as a minimum. If you are able to publish all your data in a repository, include them in the cross-linking. Also include the software image .
Minimum workflow
Linking Article, Data and Code
When reaching a milestone and/or publishing data or a paper, one should integrate the corresponding feature
branch
into the main (/development) branch. Then create a git tag to create a snapshot of the code, you are referring to and basing your results on.
Add your data from simulation to a data repository (e.g. TUDatalib) to get a PID for your data. You can use that PID to reference your data unambiguously.
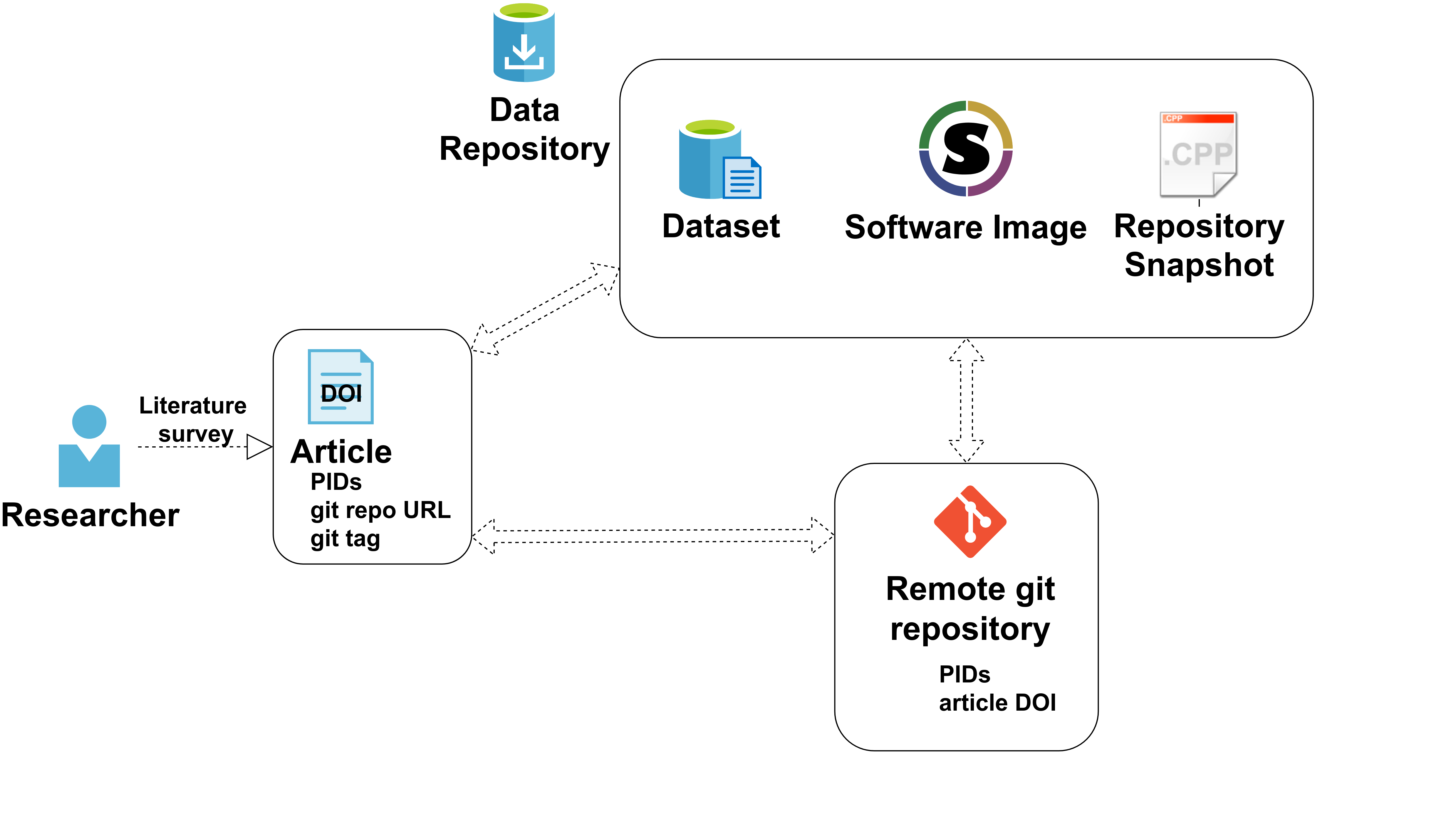
The git tag, as well as the PID of the secondary data are referenced in the publication and the publication is submitted to a repository (e.g. arXiv), which further produces the PID of the publication. The description of the git tag is updated with the PID of the secondary data archive and the publication’s PID, and the meta-data of the secondary data on the data repository are updated with the git tag and the PID of the publication.
Organization and display of data
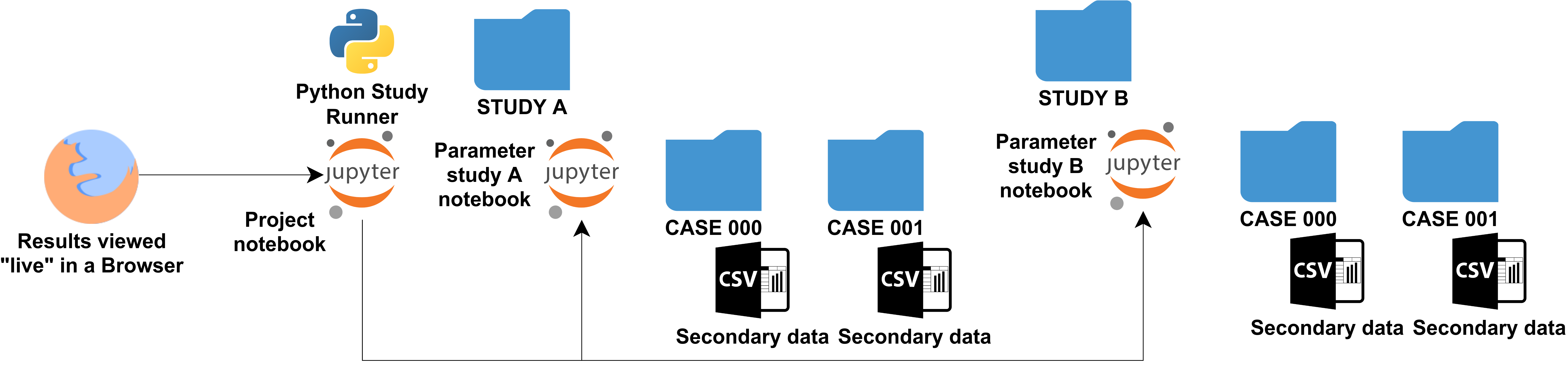
Jupyter notebooks have emerged as a convenient way of processing and documenting data, e.g., explore tables or generate plots. In our workflow, the data generated through simulation is processed in Jupyter notebooks (see also Section 2.8). In the minimum workflow, the Jupyter notebooks are additionally exported in the HTML format, within the directory structure shown in the figure above. The HTML export makes it possible to view the results without actively running a Jupyter instance, enabling quick retrospective inspection. The small secondary data (e.g. CSV files in the figure) are archived together with the Jupyter notebooks and their HTML exports, the tabular information of the parameter study that connects case IDs with parameters, and the simulation input files. In other words, the secondary data archive contains the complete directory structure from the figure above, including all parameter studies reported in a publication, but without the large simulation results.
(modified fromA Research Software Engineering Workflow for Computational Science and Engineering)
Full workflow
The full workflow cross-linking adds primary data (simulation results) and containers to the minimum workflow cross-linking. Each parameter study is stored separately because of the expected file sizes.
Where to start
To start continue with publishing with TUDatalib.
Or get a list of our articles assigned to this chapter via the tags on the top of every page.